DNA for the MCAT: Everything You Need to Know
Master MCAT DNA concepts with high-yield content, targeted practice questions, and detailed answers to boost your understanding and improve your score.
(Note: This guide is part of our MCAT Biochemistry series.)
Part 1: Introduction
Part 2: Structure of DNA
a) Nitrogenous bases
b) Phosphate backbone
c) G-C content
d) DNA synthesis
Part 3: The Central Dogma
a) Transcription
b) Translation
c) Exceptions to the central dogma
Part 4: Epigenetics and imprinting
a) Histone structure and function
b) X inactivation
Part 5: Disorders and cancers
a) Oncogenes and tumor suppressor genes
b) Chromosomal defects
Part 6: High-yield terms
Part 7: Passage-based questions and answers
Part 8: Standalone questions and answers
----
Part 1: Introduction
Understanding DNA and its function is crucial on the MCAT, especially as DNA is the biological basis of life. DNA, or deoxyribonucleic acid, is one of the two main molecules of nucleic acids (the other being RNA, or ribonucleic acid). DNA stores and selectively expresses the genetic information our cells need to function. DNA is carefully replicated and inherited from one generation of cells to the next to avoid any errors. Errors in DNA replication can vary significantly, from unnoticeable mutations to life-threatening illnesses, such as cancer. As a result, proper DNA regulation is vital to cell survival.
This guide will take a tour through the structure and function of DNA and what can go wrong. When studying DNA, it’s easy to confuse the names of the bases and building blocks of DNA as well as the different modifications DNA undergoes in the cell. It’s helpful to create a list outlining why each component or modification of DNA is necessary for its function. Throughout this guide, you will also see bolded terms that will be important to recall on the exam.
Let’s dive in!
Receive a free MCAT Question of the Day—written by a 528 scorer
Get every last bit of practice in before test day with a free MCAT question delivered straight to your inbox daily.
----
Part 2: Structure of DNA
Let’s take a close look at the structure of DNA. DNA exists as a double helix—an identifying structure that is unique to this biomolecule. In a double helix, two strands of DNA wrap around each other and form a twisted ladder pattern. There are two important parts of a DNA molecule: the nucleotides and the sugar phosphate backbone. Four unique nucleotides that bond with each other to create the rungs of the ladder: adenosine monophosphate (adenine), guanosine monophosphate (guanine), cytidine monophosphate (cytosine), and deoxythymidine monophosphate (thymine). Each nucleotide has a unique structure shown below.
Figure: Nucleotide bases used in the structure of DNA
What do each of these four nucleotides have in common? All of the bases have a very similar structure. Each nucleotide consists of a phosphate group (PO4-), a nitrogenous base, and a pentose sugar that links the other two groups together. In contrast, nucleosides are molecules that only consist of a nitrogenous base and a pentose sugar.
a) Nitrogenous bases
The nucleotides have different nitrogenous bases. There are two classes of nitrogenous bases: purines and pyrimidines. While both purines and pyrimidines are aromatic nitrogenous bases, they differ in size and ring structure. Purines are made of two rings and are larger than pyrimidines, which contain only one ring.
Take a close look at the structure of nitrogenous bases of the nucleotides. Adenine and guanine have purine bases, while cytosine and thymine have pyrimidine bases. Adenine and guanine are often broadly classified as purine nucleotides, while cytosine and thymine are classified as pyrimidine nucleotides. It’s important to remember adenine and guanine still have differences in the organic structure: for instance, in the number of attached carbonyl (C=O) bonds and where they are located. The same goes for cytosine and thymine. Memorizing each nucleotide’s structure will be crucial for test day.
Each nucleotide follows strict rules for bonding with other nucleotides. In DNA, purine bases bond with pyrimidine bases instead of other purines, and vice versa. In particular, guanine and cytosine almost exclusively bond together while adenine and thymine bond with each other. This bonding is known as Watson-Crick base pairing. Scientists have discovered exceptions to these rules, but you won’t be tested on them.
When guanine and cytosine bond, three hydrogen bonds form. When adenine and thymine bond, two hydrogen bonds form. We’ve illustrated these bonds below.
Figure: Hydrogen bonds provide stability between bonded bases and prevent rotation of bonded nucleotides.
These hydrogen bonds provide stability to the DNA, making the guanine and cytosine pair and adenine and thymine pair favorable. An easy way to remember which nucleotides bond together is by memorizing the letters AT and GC together. This will remind you that adenine (A) bonds with thymine (T), while guanine (G) bonds with cytosine (C).
b) Phosphate backbone
Imagine having a ladder with only rungs. It wouldn’t function as a ladder at all! A ladder needs its side rails to work. Similarly, DNA needs a backbone to hold the double helix structure together.
The DNA backbone is known as a sugar phosphate backbone. The sugar phosphate backbone consists of a repeating pattern of sugar and phosphate groups bonded together.
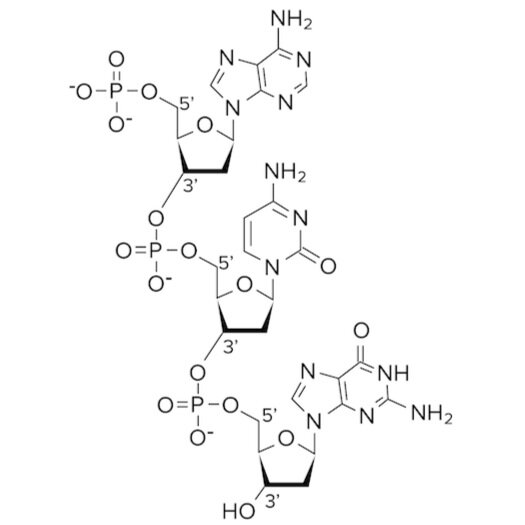
Recall the structure of a nucleotide and that each nucleotide base contains a phosphate group and pentose sugar. The phosphate group is attached to the 5’ carbon of the pentose sugar. In the sugar phosphate backbone, the phosphate group of one nucleotide creates a phosphodiester bond with the 3’ carbon of the pentose sugar on the adjacent nucleotide. Try to trace the locations of the phosphodiester bond on the diagram below.
Figure: Phosphodiester bonds in the sugar phosphate backbone.
Phosphodiester bonds exist between adjacent nucleotides to create the backbone of DNA. An important fact to know about DNA is that its backbone is negatively charged. Can you see why? The backbone is negatively charged because the phosphate groups carry charged oxygen atoms. This negatively charged backbone creates an attractive force between the aqueous, polar environment and the DNA molecule.
Consider what might happen if the negatively charged backbone were on the interior of the molecule and the aromatic bases were on the exterior of the molecule. This would lead to a very unfavorable energy conformation—the two negative charges on either side of the backbone would repel each other, and the aromatic bases would not be soluble in water at all!
c) G-C content
One of the most important ways to analyze the nucleotide composition of DNA is to calculate its G-C content. G-C content is a measure of the percentage of nucleotide bases that contain guanine or cytosine in a fragment of DNA. Why might this percentage be important?
Recall from our discussion of guanine and cytosine bonds that G-C bonds are more stable than A-T bonds. When guanine and cytosine bond, three strong hydrogen bonds are created. Breaking a single hydrogen bond takes a significant amount of energy, let alone three.
G-C content is important because it determines the melting point of DNA, as well as its accessibility by polymerases. A higher G-C content means that there are a greater number of guanine-cytosine base pairs holding the two DNA strands together. This means there are also more hydrogen bonds. A greater amount of energy is needed to dissociate the two strands, leading to a higher melting point. A lower G-C content means the opposite. Less energy is needed to dissociate the DNA strands, lowering the melting point and making the DNA more accessible to polymerases.
The MCAT may ask you to calculate nucleotide composition using Chargaff’s rules. Chargaff’s rules state that the ratio of purine nucleotides to pyrimidine nucleotides in DNA is 1-to-1. In fact, the ratio of guanine nucleotides to cytosine nucleotides and the ratio of adenine nucleotides to thymine nucleotides are also each 1-to-1.
Chargaff’s rules hold true for any piece of DNA we come across. Why? Recall that adenine and thymine bond together while guanine and cytosine bond together. Since these bases always bond in this pattern—and DNA consists of only bonded nucleotides—there must be a 1-to-1 ratio of adenine to thymine and guanine to cytosine. To understand the first part of Chargaff’s rules, let’s look at an example.
Consider the DNA strand pictured below. If we count the number of purine and pyrimidine nucleotides, we get a 1 to 1 ratio. If we count all of the nucleotides, we get a 1-to-1 ratio of guanine to cytosine and a 1-to-1 ratio of adenine to thymine.
Figure: Every DNA strand follows Chargaff’s rules.
Now, let’s imagine a fragment of DNA had a G-C content of 30%. The low G-C content tells us that this piece of DNA has a low melting point and is more accessible to polymerases. Chargaff’s rules allow us to determine that the DNA must be 15% guanine and 15% cytosine.
We also know that the rest of the nucleotide content in the DNA must be composed of adenine and thymine (since DNA has two types of nucleotide bonds). Thus, the percentage of combined adenine and thymine content must be 70%, or 35% each.
We now know our DNA fragment consists of 15% guanine, 15% cytosine, 35% adenine, and 35% thymine. If we add the percentages of the purine nucleotides together and pyrimidine nucleotides together like in Chargaff’s rule, we get 50% purine content and 50% pyrimidine content: resulting in a 1-to-1 ratio, just as Chargaff described.
Gain instant access to the most digestible and comprehensive MCAT content resources available. 60+ guides covering every content area. Subscribe today to lock in the current investments, which will be increasing in the future for new subscribers.
d) DNA synthesis
As cells grow and divide, they also need to replicate their DNA. How are they able to exactly duplicate these lengthy sequences of nucleotide bases?
First, we need to understand the directionality of DNA. Each end of DNA is assigned a number, 5’ or 3’, based on the orientation of pentose sugar in the nucleotides. The 5’ end of DNA refers to the end of the backbone chain where the phosphate group is bound to the 5’ carbon of the pentose sugar. The 3’ end of DNA refers to the end where the 3’ carbon creates a phosphodiester bond with the adjacent nucleotide.
When DNA bonds together, the two strands run in opposite directions or (antiparallel). One strand of DNA runs in the 5’ to 3’ direction, while its complement runs in the 3’ to 5’ direction. (It may be helpful to refer to the previous image to see how this fits together.)
Duplicating DNA requires that the helix “unzip” momentarily so its nucleotides can be read. Because single-stranded DNA is unstable and prone to degradation by DNA nucleases, DNA unzips in small intervals. DNA replication starts at the origin of replication, a sequence rich in adenine-thymine bonds. Chromosomes of eukaryotic organisms may have multiple origins of replication, thus allowing replication to occur simultaneously at multiple different sites.
Two important enzymes, helicase and DNA topoisomerase, begin to unzip the DNA and relax the coiling in the DNA, respectively. (As DNA is unwound, it can form tangles known as supercoils. Topoisomerases help to unwind the tangled coils that begin to form by making selective cuts in the phosphate backbone and repairing them.) The unzipping progresses in both directions away from the origin of replication, so replication can progress in both directions and decrease the amount of time needed.
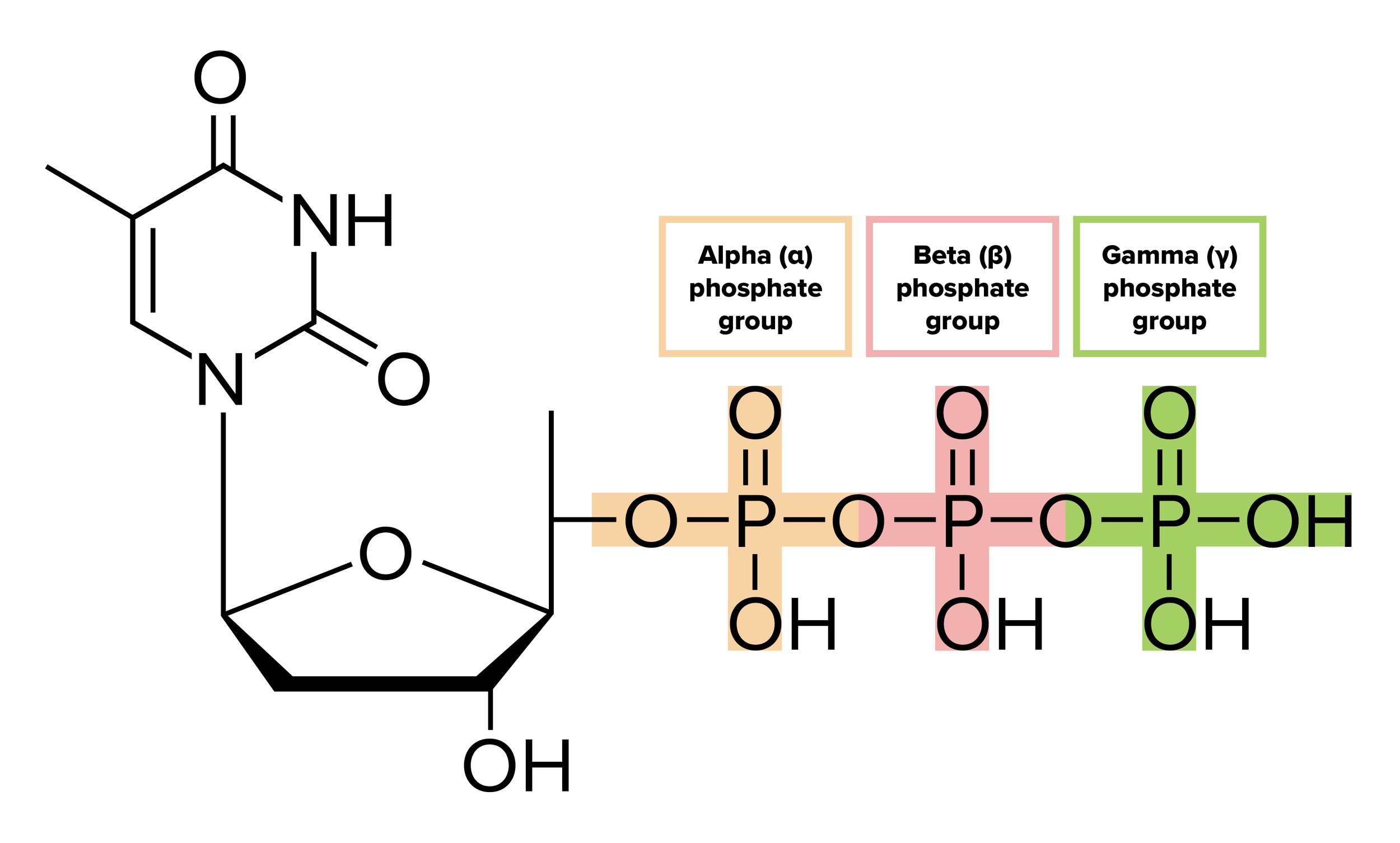
DNA polymerase (sometimes called DNA pol) can continuously add nucleotides triphosphates to create a new daughter strand. It’s important to note that the nucleotides added by DNA polymerase are in the form of nucleoside triphosphates (NTPs). Each phosphate group is designated alpha (α), beta (β), or gamma (γ) depending on its distance from the nucleoside. When a nucleotide is incorporated into the growing DNA strand, the β and γ phosphate groups are cleaved off, releasing the energy that drives the formation of the phosphodiester bond. In fact, when we think of all the instances where a high-energy triphosphate like ATP is hydrolyzed to fuel a reaction, it is generally the γ phosphate group that is cleaved off, leaving behind ADP!
Figure: Deoxythymidine Triphosphate, the nucleotide form of thymine prior to its addition by DNA polymerase. The beta and gamma phosphates are cleaved during this addition.
DNA polymerase creates a translated strand that is complementary. The translated (or new) strand will contain an adenine base (A) at every position there is a thymine base (T) in the DNA sequence, a guanine base (G) at every position there is a cytosine base (C) in the DNA sequence, and vice versa.
DNA polymerase synthesizes DNA but with a catch. The polymerase only creates DNA in a 5’ to 3’ fashion. That means the template strand the polymerase is attached to must run in the 3’ to 5’ direction. While this is the case for one of the strands (called the leading strand), recall that the two strands of DNA are antiparallel—so the other one (called the lagging strand) runs in the 5’ to 3’ direction.
DNA replication of the leading parent strand runs smoothly. Replication of the lagging parent strand is discontinuous. Because DNA polymerase only runs in the 5’ to 3’ direction, DNA primase continuously adds new primers along the lagging strand. DNA polymerase uses these primers to create short sequences of DNA known as Okazaki fragments. At the end, DNA ligase seals the fragments together and fills in any gaps that may have been left behind to create a new daughter strand.
Figure: DNA replication.
DNA is replicated in a semiconservative manner. This means that during replication, each strand acts as a template, or parent strand, for a new DNA molecule. After one round of DNA replication and mitosis, each daughter cell contains one new DNA molecule composed of one strand of original DNA (from the parent cell) and one newly synthesized strand of DNA.
When DNA is being replicated, DNA polymerase also proofreads the new daughter strand being synthesized. The polymerase will ensure that the complementary nucleotide is paired to the correct nucleotide on the parent strand of DNA.
Many of the mistakes that DNA polymerase makes are quickly resolved through proofreading. For mistakes that aren’t caught during proofreading, the cell relies on mismatch repair. Mismatch repair occurs in the G2 phase of the cell cycle to correct any leftover errors from synthesis. Enzymes such as MSH2 and MLH1 are responsible for detecting, removing, and replacing incorrectly paired nucleotides.
As the replication of DNA is repeated multiple times, the ends of the DNA molecules become shorter and shorter. These are regions known as telomeres. Telomeres protect and stabilize the coding regions of DNA. As telomeres shorten, they eventually reach the shortest allowable length, and that cell stops replication. The Hayflick limit refers to the number of cell divisions at which this point is reached.
----
Gain instant access to the most digestible and comprehensive MCAT content resources available. 60+ guides covering every content area. Subscribe today to lock in the current investments, which will be increasing in the future for new subscribers.
Not ready to commit to your MCAT prep at this time?
Enter your name and email for a free MCAT question delivered straight to your inbox daily.